Spatial genomics is an emerging technology that provides spatial resolution to RNA-Seq data. It combines the state-of-the-art imaging processing of tissue sections and well-established high-throughput transcriptomic quantification to enable the discovery, identification, and/or study of cell types and gene expressions for a given physiological region. It has not been used for long, but the research community has shown great interest and confidence in it.
Currently, a few commercial platforms are available for spatial genomics research, and the University of Minnesota Genomics Center and University Imaging Center are working hard to bring some of the technologies to campus, including the CARTANA system (now part of 10x genomics). MSI’s Research Informatics Solutions group is facilitating this process by bench-marking the sensitivity and specificity of publicly available datasets.
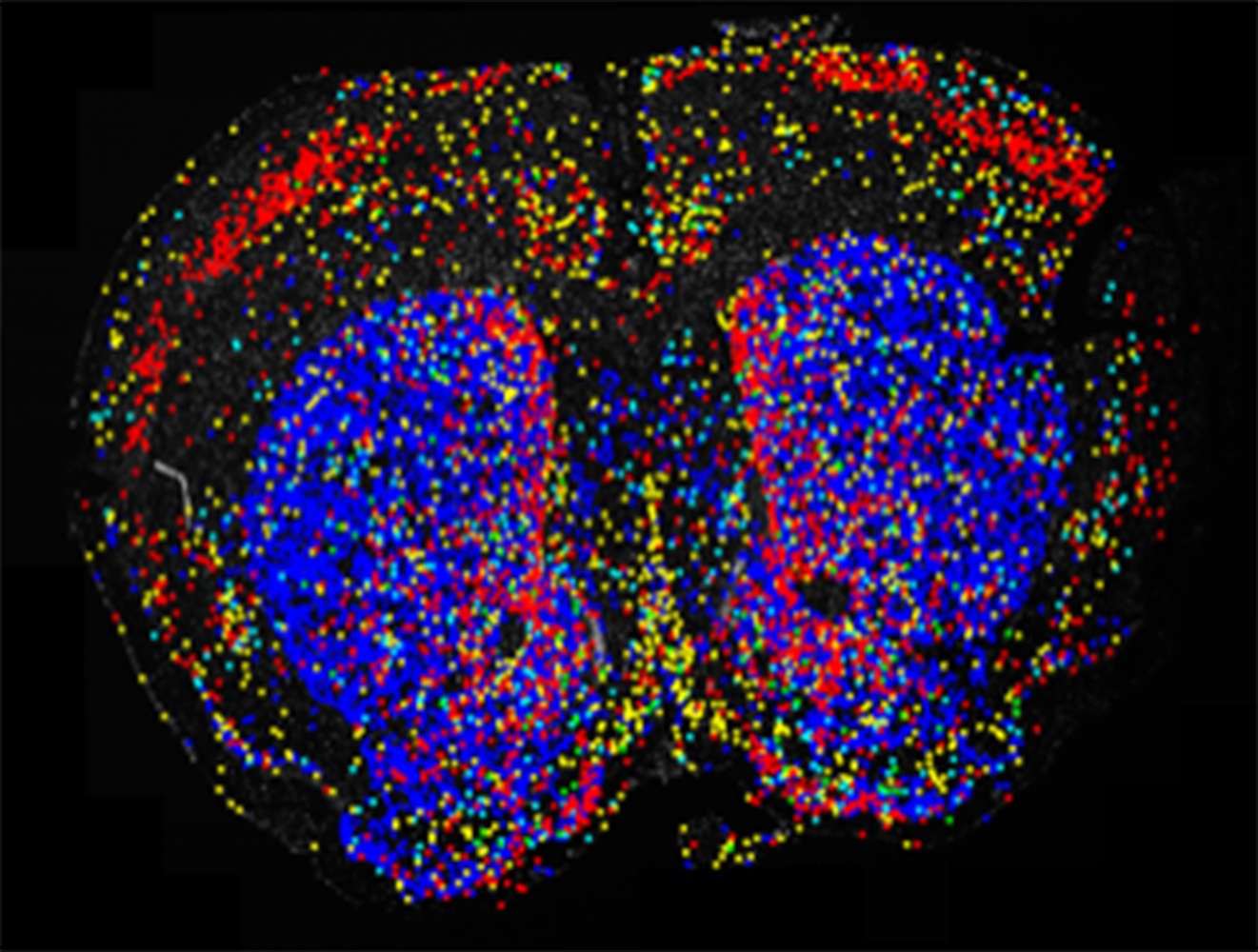
In collaboration with MSI PIs Patrick Rothwell (associate professor, Neuroscience) and Walter Low (professor, Neurosurgery), Dr. Ying Zhang (MSI) and Dr. Thomas Pengo (University of Minnesota Informatics Institute) are using MSI’s high-memory computing nodes to process the gigantic image dataset. The initial result has been posted at the GitHub UMN site for the use of interested U of M research groups, and one processed image is shown here. The image uses a public dataset from CARTANA that was processed locally.
In the image below, a few marker genes were probed for expression in a slice of mouse brain with visible structure of striatum, the critical component of the motor and reward systems and key player in drug addiction. The colored dots are cells expressing the specific marker genes. Blue is Penk-expressing D2-MSN neurons, the dominant cell types (95%) in human striatum. Yellow is striatal fast-spiking interneurons. Cyan is striatal low-threshold spiking interneurons. Red is the supportive astrocytes distributed in whole brain. There are very few green dots that represent Slc17a8-expressing cholinergic interneurons.
Research Computing partners:
- Minnesota Supercomputing Institute
- University of Minnesota Informatics Institute